
Marcin Czerwinski on Twitter: "DEL phenotype attributed to RHD Exon 9 sequence deletion: slipped-strand mispairing as probable culprit #Rh #blood_group https://t.co/BHRIJBmQNk https://t.co/TrFileAu9A" / Twitter
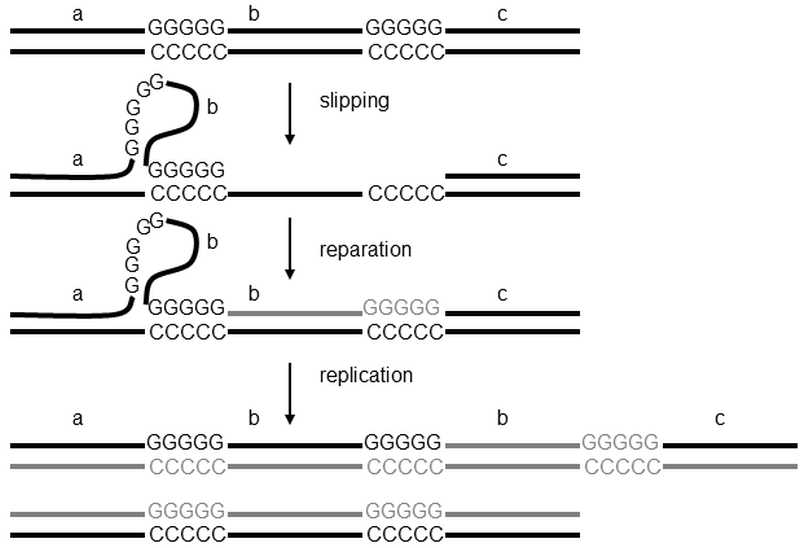
VI.2.2.4 Multiplication can also be the product of the nucleotide slipped�strand mispairing mechanism. | Frozen Evolution. Or, that's not the way it is, Mr. Darwin. A Farewell to Selfish Gene.
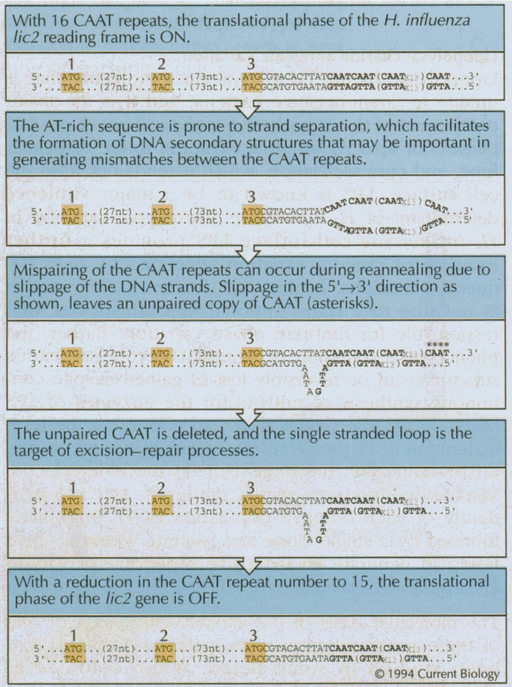
Chapter 23 Notes: Evolution of Genetic Systems: Figure WN23.4 - Possible mutational mechanism responsible for LPS phase variation in H. influenzae, based on slipped-strand mispairing (Levinson and Gutman 1987; Weiser et al. 1989).

Figure 2 from Common strategies for antigenic variation by bacterial, fungal and protozoan pathogens | Semantic Scholar

Slipped Misalignment Mechanisms of Deletion Formation: In Vivo Susceptibility to Nucleases | Journal of Bacteriology

A DEL phenotype attributed to RHD Exon 9 sequence deletion: slipped‐strand mispairing and blood group polymorphisms - Lopez - 2018 - Transfusion - Wiley Online Library

A model for slipped strand mispairing during the PCR showing why n–2... | Download Scientific Diagram

A Computational Method to Quantify the Effects of Slipped Strand Mispairing on Bacterial Tetranucleotide Repeats | Scientific Reports



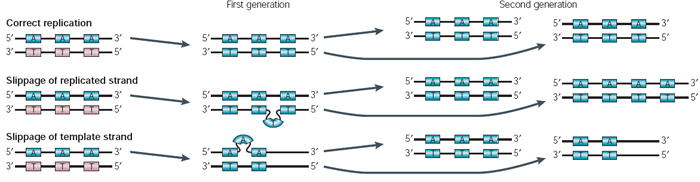
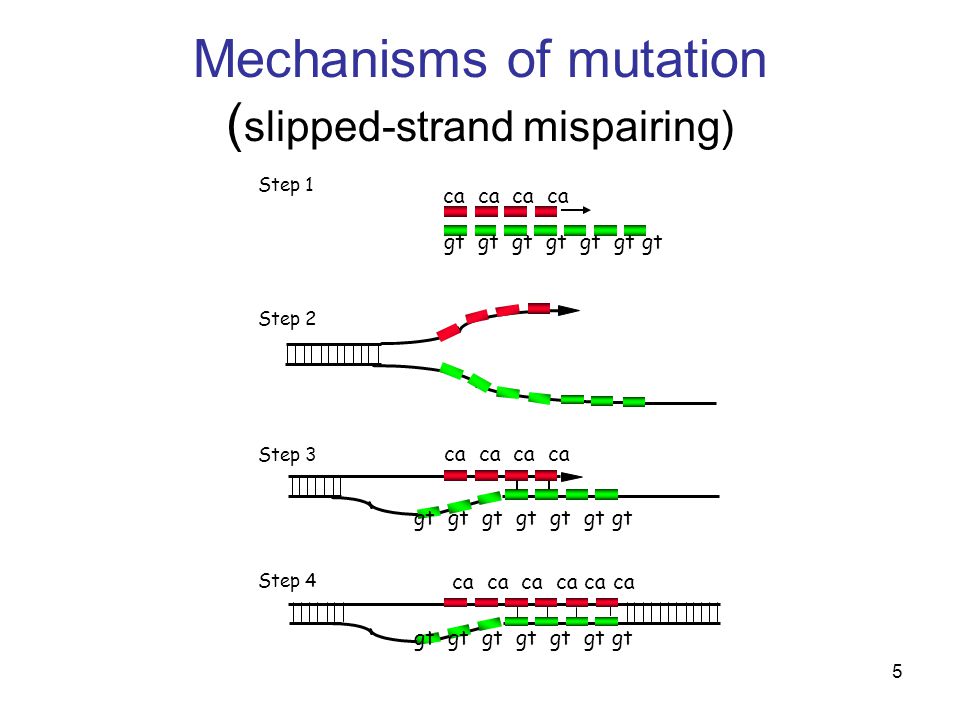

![PDF] Slipped-strand mispairing: a major mechanism for DNA sequence evolution. | Semantic Scholar PDF] Slipped-strand mispairing: a major mechanism for DNA sequence evolution. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/b51bdbdd1645f35e6c13ab4591ffba81451dc2d1/6-Figure3-1.png)





